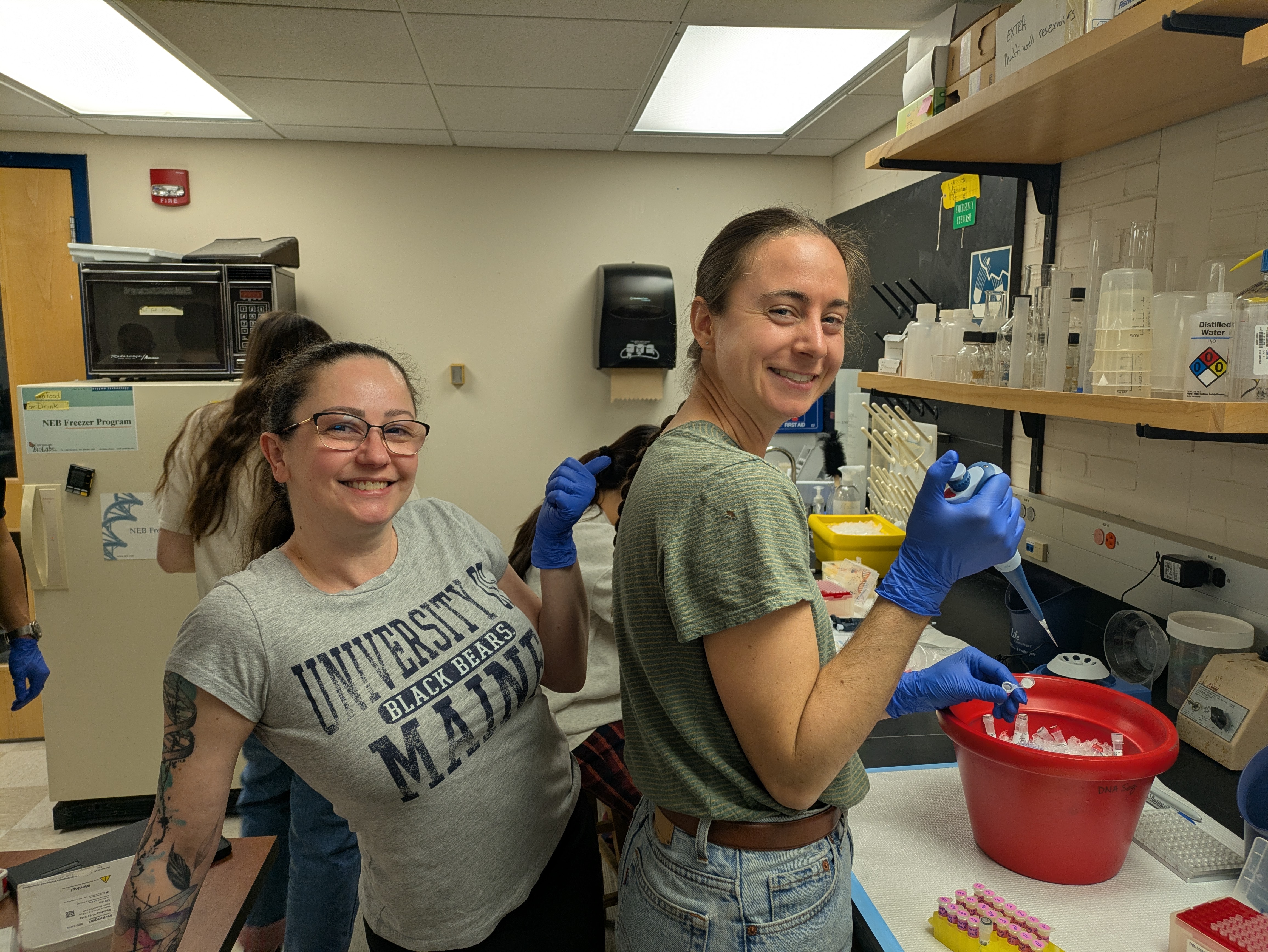
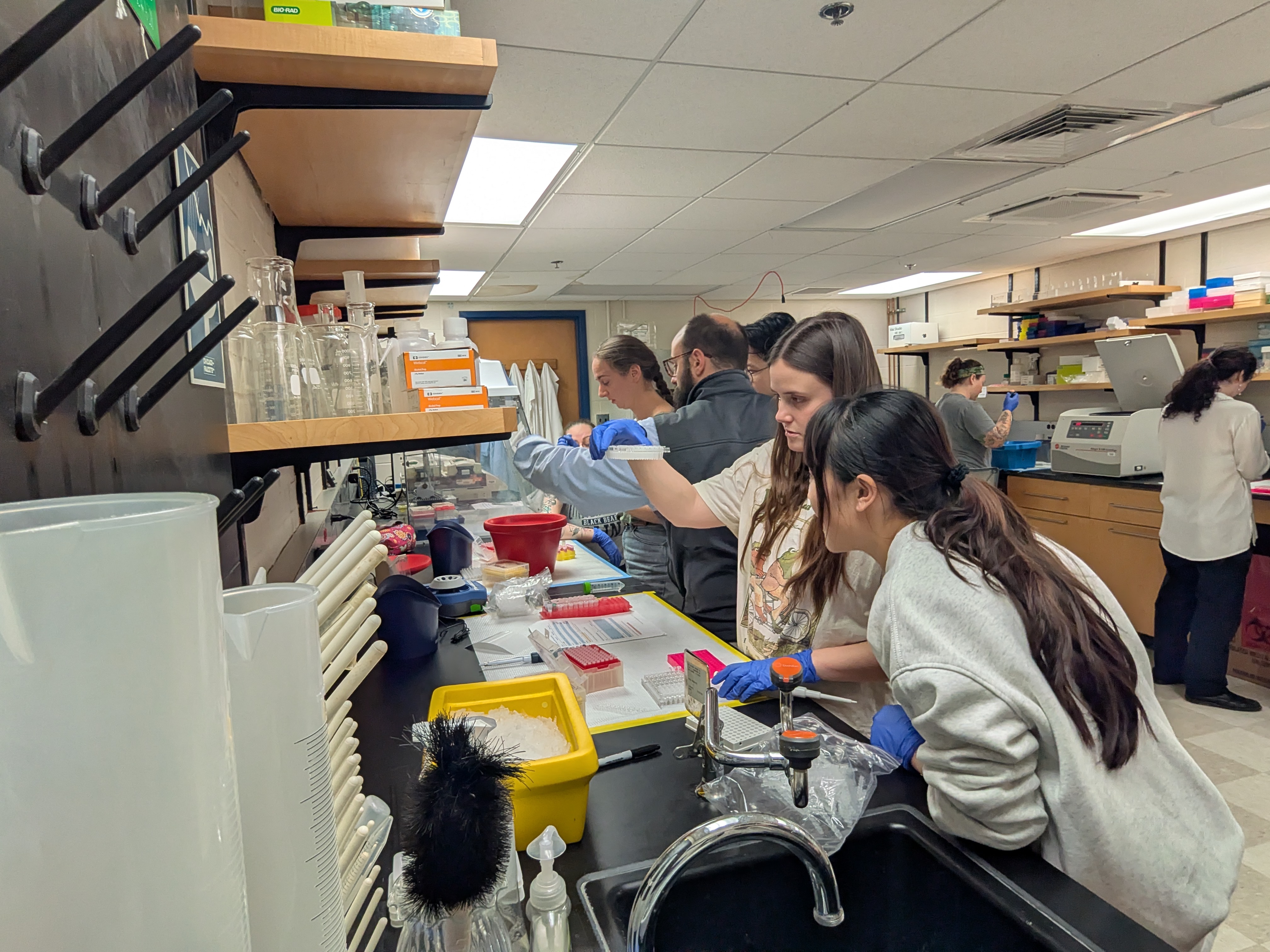
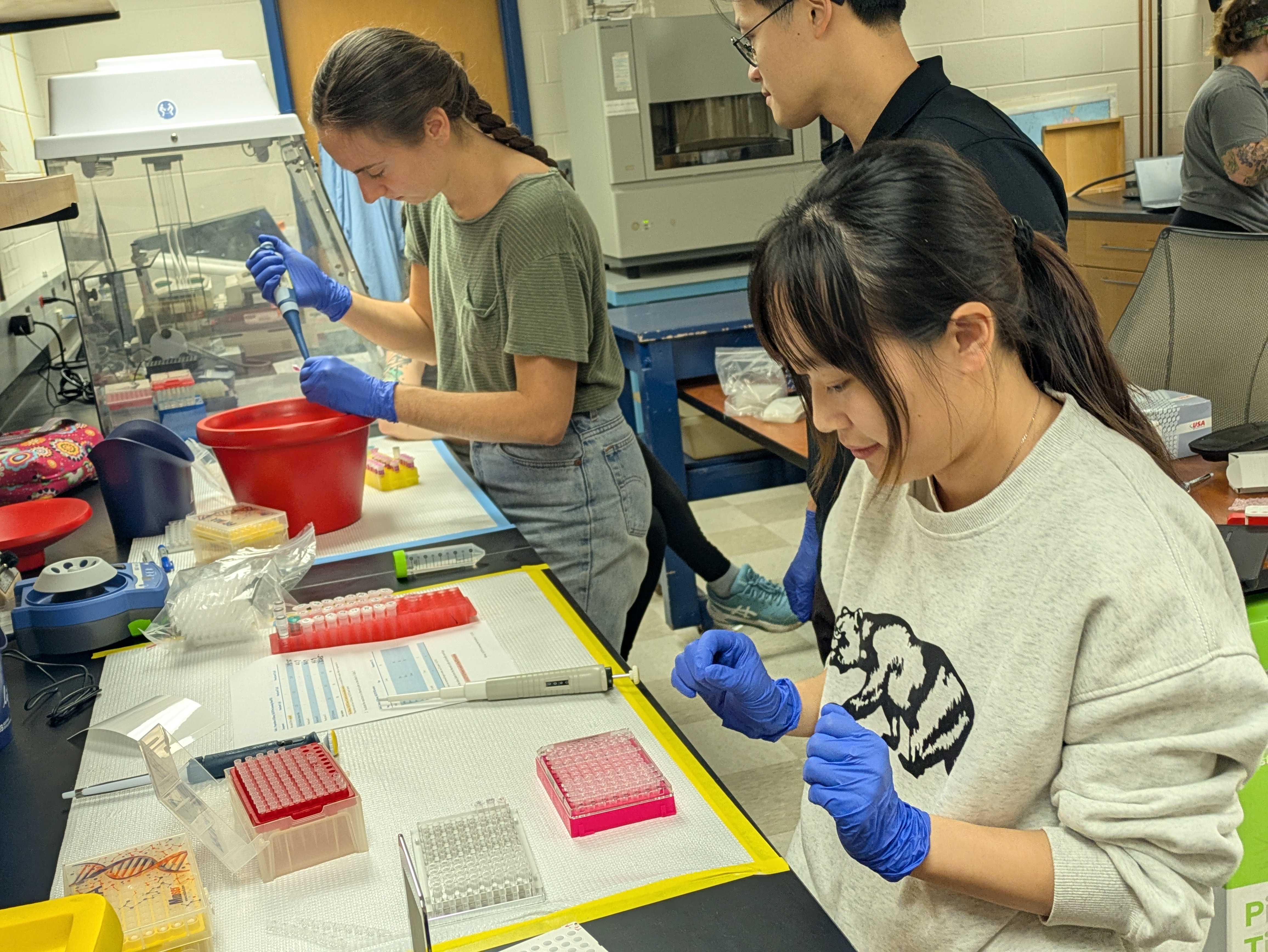
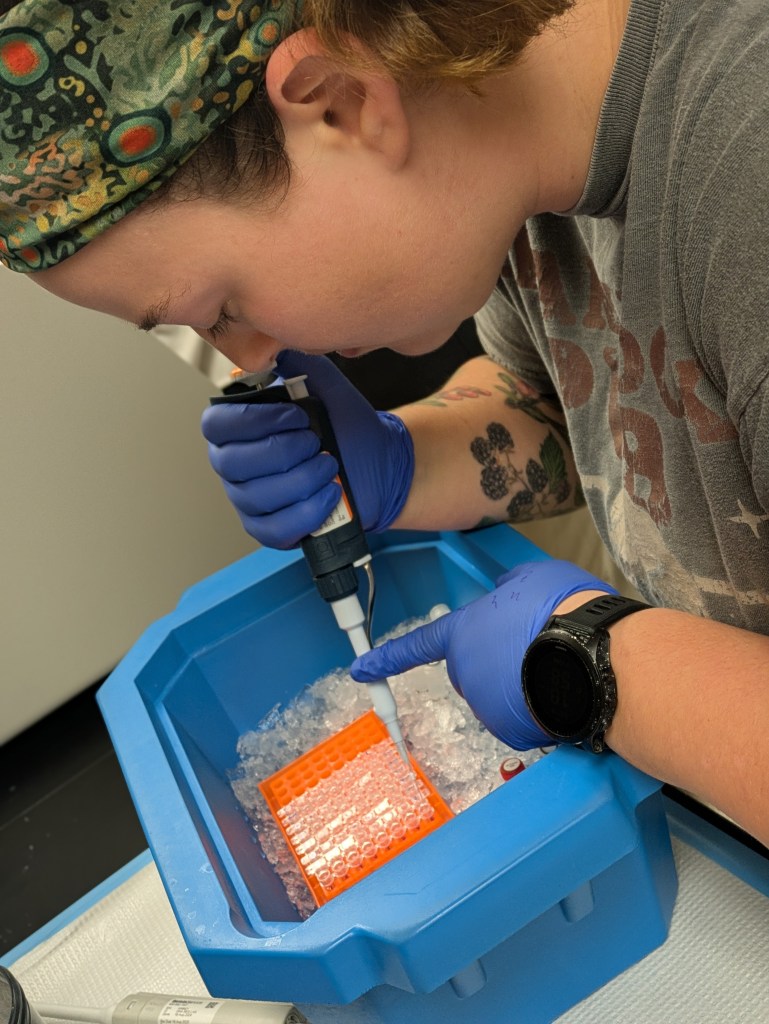
We’ve all felt the thrill of synchronicity when meeting someone for the first time and realizing how we have much in common, but when this occurs for a dozen people simultaneously, who go on to share ideas and excitement for 13 hours straight, it’s magic. Thus, Professor of Anthropology, Roberta Raffeta’, created magic when she invited a group of microbiome and health science researchers together for the “Integrating Qualitative and Quantitative Data in Microbiome Research and Postgenomics: Toward an Interdisciplinary Dialogue” workshop at the Università Ca’ Foscari Venezia. Roberta works with many disciplines, including microbiome, computational, health, and social sciences, and her work often focuses on how research is designed, implemented, and interpreted. Her work across disciplines gives a larger view on how different disciplines approach similar research, as well as provides her with a rich network of colleagues.

The workshop was in the spirit of a project that Roberta, Prof. Nicola Segata, Prof. Elena Bougleux, and others investigating the sharing of microbes at Antarctic stations based on social interactions and shared spaces. That project is a clear example of how social systems can determine who you interact with, where, and how, and thus which microbes might get shared between you or between everyone at the base. While those data were still being processed and not shared, we did get to hear a bit about the journey to Antarctica from Elena.
After introducing the project, workshop, and ourselves, we began with a presentation by Professor Federico Russo. Her talk focused on how health is quantified, and how that alters the way we design and perform health research. Health is a complex concept that has biological, chemical, physical, and social aspects which can be measured and metric-ified. For example, there are bio-logical versus bio-social metrics of health (social effects of disease, feelings about health and outlook), and everything in between which can be measured to assess the state of health or disease. In addition, health research uses the term “social determinants”, which is similar to what biologists, microbiologists, or ecologists mean when using the term “environmental factors”, but these often refer to the same thing. These factors are the collection of host, social/community, environmental, and geological features which affect who you are and what you encounter in your life. Some of them seem very specific, like age, but the expression of age can be modified by staying active, eating well, managing stress, and avoiding pollution, so on its own, knowing someone’s age might not be useful information. Thus, to study health, we use both markers, like age, to tell us potential outcomes or indicators of one’s health state, and social determinants, like lifestyle features, to tell us possible causes or mitigating factors of health.
Yet, one cannot slice the concept of health into a thousand measurable factors and then expect to re-assemble them back into the concept it was – because what makes us feel well or healthy is not necessarily having or knowing that our biological metrics are good, it’s feeling well even if our biological markers are out “normal” parameters or when, on paper, we are sick. This brings up a concept that I had discussions about within this workshop group, with the faculty at Ghent, and with researchers through MSE and MiSt: that your social factors and the support network around you strongly influence whether you feel well or unwell, regardless of what your biological markers would suggest.
There is also a focus in healthcare on regaining health when someone is sick, with social or institutional support system for that (rehabilitation clinics, etc.), but there is not always an institutional focus on understanding how people stay healthy, in part because this is seen as a personal choice and not a result of adequate access to public resources (fresh food, water, air, shelter, education, safety), or as a function of useful public health policies which make is easy for people to take care of themselves. Simple features like sidewalks, bike paths, local grocery stores, free public restrooms, shade and places to sit, are all features that allow people to stay active, get around, stay healthy, and use their public spaces.

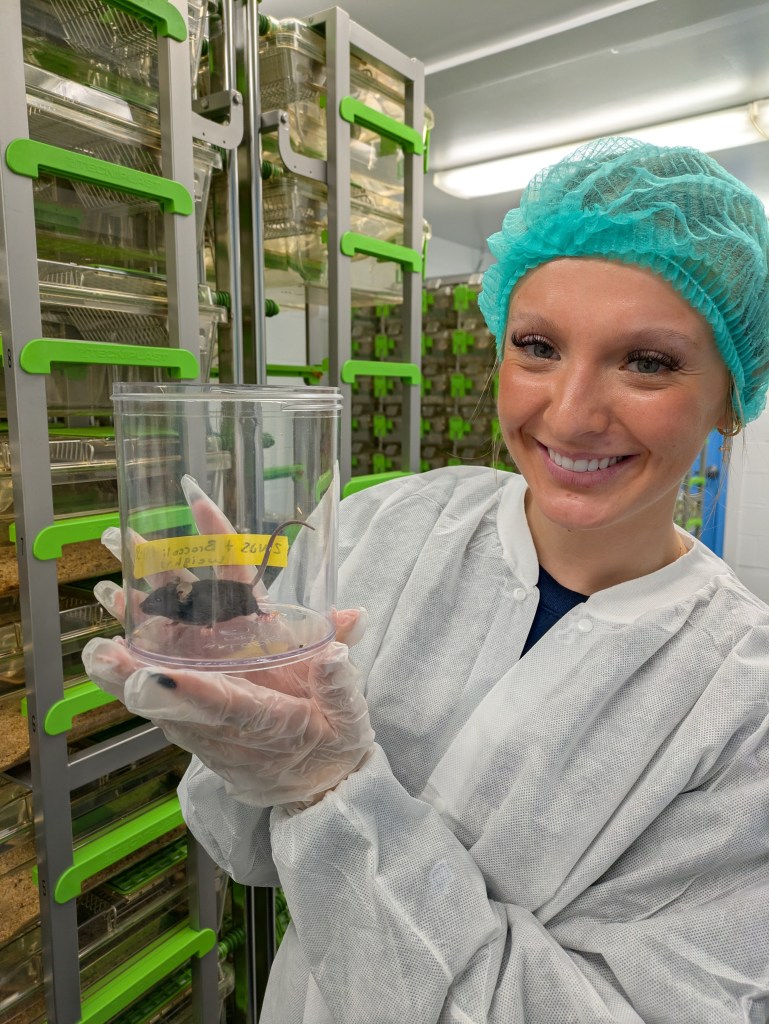
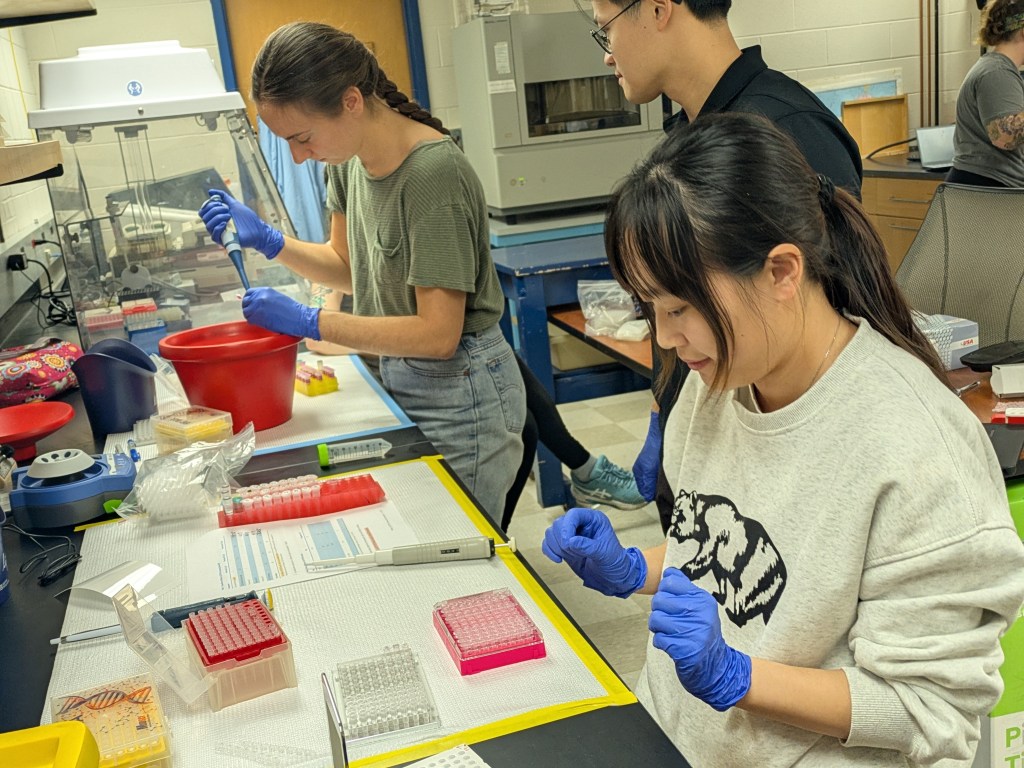
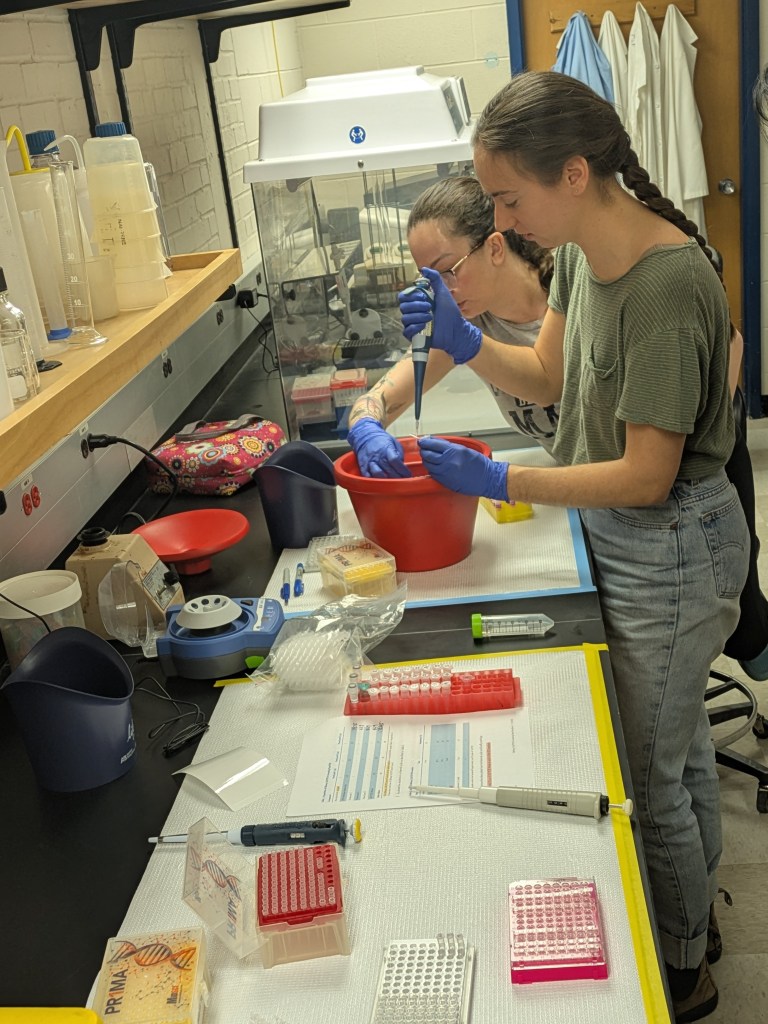
My research talk was next, and I focused on the steamed broccoli sprout intervention trial I completed a few years ago with Yanyan Li. That was a pilot study, which recruited 20 people to steam and eat broccoli sprouts every day for a month, to measure any changes in the gut microbiome, the metabolites it was producing, and whether gut bacteria would convert the inactive glucoraphanin in sprouts into the anti-inflammatory compound sulforaphane. Rather than focus on the microbiological, metabolomic, or diet survey results, I presented everything which went unexpectedly in the study, and what people told us about the challenges to consuming daily sprouts. This, in fact, was the real goal of the study, to understand which aspects of the diet would be challenging, or rewarding, and to try and make things as easy as possible. My observations on the diet study sparked excellent discussion, which gave me plenty of ideas on framing the scientific manuscript and what we learned from our participant’s data, as well as a perspective piece on the design and implementation of the study and what we learned from our participants’ feedback (not their data).
Another study observation which was also a reoccurring workshop discussion was the need for health studies that start with people’s perspectives and patient’s identification of problems, which then work backwards to understand how the microbiome is involved. This style of research is case-study and health engineering research to test applied research questions, and is needed in addition to the large-scale, double-blind experiments to test basic research questions. We talked at length about how most large-scale diet surveys are inadequate, no matter how detailed they are, because they become so vague as to be useless when they are generalized to ask about all possible food item diets. Most diet surveys that are meant to be broad ask for too much detail about things which are considered superfluous for individual research projects, and too little detail on critical info. For example, most diet surveys that gather diet history (eating habits over the last 6 – 12 months) are underpowered to assess fermentation products, don’t ask how people cook and make decisions about diet, and are quantified to assess compliance to an idealized and single idea of a healthy diet, even if it doesn’t work for every person (ex. dairy is good on many diet surveys but there is no place to select that you don’t consume dairy because you are allergic to it).
This discussion carried across lunch, during which we diverged into many animated conversations only to bring it back to quality of information in the presentation after lunch, by Professor Lisa Lehner, a health researcher who presented reflections on three research projects, in which the availability and completeness of data about patients was lacking, and this stymied researchers’ attempts to understand public health for three disease models, such as how these diseases are transmitted, how people access health care, and how migration, homelessness, or simply traveling often can impact access to care. For example, in trying to study human papilloma virus cases, her research found that country of sampling was included in patient records but not country of infection. Similarly, the idea of “where do you live”, “where are you from”, or “where have you been recently where you might have contracted this infection” are very different questions with different contexts to the answers, not all of which will provide useful information for this specific study depending on which the patient was asked and what their history was.
Information sharing is both the key and the challenge, and we discussed how often the information to resolve public health crises exists, but it’s not all in one place, it’s missing information, it doesn’t contain the right context, institutions don’t want to share, some information is private or access is limited even to medical providers, some information is not retained, and even when we get much of this in one place the amount of detail is overwhelming to the point where researchers need to spend years trying to figure out how to make it useful – what’s important to know? What is a ‘red herring’? — before it even gets to be used for infection tracking, treatment, and prevention. Her research also highlighted inequities in healthcare, and that sometimes information that would help us understand infection data can’t be made public, because you might reveal information about sensitive populations that can be used for discrimination. For example, people without health insurance might be migrants that are no longer on active visas, and quantifying how many people can make them a political target.
The discussion after the talk was so engaging, and blended into the focus of another speaker, so we informally heard from Professor Donato Giovannelli, who was not able to present his talk (yet!) because the workshop was running later than expected (we all had so much to talk about!) and he had to catch a train. Donato is a geomicrobiologist examining microbes in extreme environments to understand how they survive and function. He has a particular focus on the microbial fixing of gaseous hydrogen which produces water, and how biological sinks for freshwater (like us walking, talking, water bottles), actually allow for the preservation of large quantities of water on Earth. Donato’s short version of his talk was focused on the scale of standardization in research. He argued that what works in the lab or the field changes based on the needs and circumstances of each project, so you need to thoroughly describe what you did but that doesn’t mean every lab uses, or has to use, the exact same protocols, kits, or methods every time. In fact, even if it were possible to replicate circumstances exactly, trying to do everything identically will just make all studies biased in the same ways. There is no way to replicate some experiments, especially with humans, because no human is ever replicable, even to ourselves.

Highlighting the importance of methods and process in research brought up another challenge in research: it is important to include vocabulary to describe what you mean, in addition to what you did, but Methods sections are often compressed by scientific journal word limits, and all the nuance and context or the problem-solving portion, gets cut for space and is over compressed to the point of being useless. Donato argued, as many scientists do, that the explanation of the methods should be the majority of the paper, and many labs do publish the long form of the methods as a co-published paper, or as a protocol paper that gets cited in the whole scientific experiment paper. Our workshop heartily agreed. Science is a process, as scientists we are creating this process and this is our most valuable contribution, more so, even, than the results.
Sometimes the drive for standardization also leaves out people who contributed to the study in a way, but their contribution is considered “not science”. For example, technicians, field personnel, or anyone who helped you access or collect samples but did not work on the samples or contribute “conceptual framing or interpretation” don’t count as authors under most journal guidelines, even if you could not have done the project without them. Similarly, if you designed your project based on feedback and direction from the public or specific audiences, this is somehow not considered “conceptualization” or experimental design (???!!!). It is always up to the author team to decide who is and isn’t an author, but without more flexible guidelines on authorship, it can be easy to remove someone’s contribution to a publication.
This type of value judgement in science segued us to our last presentation of the day, on how the terms/definitions we use, the factors we choose to study, the things we prioritize in our research, all reflect values we have placed on some things over other. It can also, unintentionally and intentionally, place inherent value on having certain study results or outcomes. And when the study is about people, placing value judgements on biological, microbiological, or social factors can create a hierarchy – you can see where I am going with this – in which certain people are implied to have superior metrics. This is something a research team and I published on in microbiome research, and has been the focus of a multi-year project led by Professor Abigail Nieves Delgado, Dr. Aline Potiron, and others, on how microbiome scientists define terms which are used to describe people, and how those definitions carry assumptions that may influence the interpretation of the results by researchers or readers.
Population descriptors are ways of, well, describing populations. However, the selection of terms, how you define them, how you categorize those terms, are often too strict to be applied to real-life people or in different communities. Using the same example of age, how do you compare age in two populations with wildly different exercise, diet, or stress load which all impact the physical or behavioral expression of age? More to the point of this talk, how do you categorize ancestry without nuance? Do you focus on genomics, or culture, or both? Do you place people in categories based on certain factors, or let them choose their own? How do you compare people in one location to another without considering the history of war, colonization, and slavery had on the ability of people to choose with whom to have children? And does ancestry even matter when you consider the myriad changes to health and the microbiome effected by what you eat, what animals and people you interact with, and where you live? Scientists all working on the human microbiome had wildly different understanding or definitions of population descriptors, and beliefs around what population descriptors were caused by or caused changes to the microbiome. And this, despite most microbiome studies having little to no need for information on ancestry or race to answer their research question.
Studies don’t gather data on every possible factor in people, that would require a detailed biography of every participant. Researchers choose what type of data to collect about someone based on what is important to the study and answering the research question – but as researchers we tend to think within the constraints of our own discipline – what info do I need for my statistics or to publish in my field? When that happens, we compress the variability of the people we study into fewer dimensions, example like taking an in-person experience at the beach and representing it as the words “seashells”, “gritty floor”, and “wet”.
Finally, a theme that came up several times during the workshop was that human social structure or institutions sometimes have so many constraints as to fail in their mission to provide resources to the public. As a perfect example, we did not have air conditioning at the university, even though was 85 degrees outside and between 90 and 105 degrees F inside some of the rooms after we had been in them for a while, because it is university policy to turn air conditioning on for the whole campus only after a certain date regardless of what is going on in real life. I don’t want to pick on the university, because this is a very common policy at universities, including UMaine. But it highlights the point that solutions (to air conditioning, microbiomes, health, etc.) need to be nimble enough to respond to local conditions using the input of people who are actually living through those conditions. And in this way, we have the crux of the workshop: how do we perform local-conscious and nimble science that is driven by the needs of actual living things or ecosystems, against the tide of institutional habit? For me, that’s going to involve more community-based and person-driven research questions, more collaboration with this remarkable group of researchers, and hopefully a lot more fresh, Italian mozzarella.