The 2nd Northern New England Microbiome Symposium
Whether we are interested in the way coastal wetlands sequester carbon, how diet affects us differently, or how public and environmental health are inextricably linked – research on the microbiome can reveal how systems connect. Join us for two days to learn about how microbial communities impact ecosystems, food production, health, and more; to hear from experts researching these issues in New England and beyond; and discuss the technology and data analysis which can boost your own research.
UMaine Buchanan Alumni House, 160 College Ave, Orono, ME 04473
If you would like to help sponsor this event, please contact sue.ishaq@maine.edu.
Agenda for June 11, 2026
8:00 – 9:00 am Breakfast, provided
9:00 – 9:15 Welcome and Opening Remarks
Dr. Sue Ishaq, PhD, Associate Professor of Microbiomes, Associate Director of Special Projects in the School of Food and Agriculture at the UMaine; member of the Board of Directors at ASM; Founder and Lead of the Microbes and Social Equity working group.
9:15 – 10:15 Plenary Presentation
“The second messenger c-di-GMP in Bordetella”
Dr. Federico Sisti, Ph.D., Investigator, Institute of Biotechnology and Molecular Biology (IBBM) of the National Scientific and Technical Research Council (CONICET). Dr. Sisti is incoming President-Elect at the American Society of Microbiology, and the first ASM President representing its global membership.

10:15 – 10:30 Break
10:30 – 12:00 Session 1: Microbial mayhem or tiny partners in health?
“Leveraging organ-on-chip technology to study gut microbiome effects on human health and disease“, Dr. Dani Brassino, PhD, Assistant Professor of Gut Microbiome and Cancer Interactions, The University of Vermont College of Medicine.
Following a B.S. in Nutrition Science and at the University of Texas, Dani Brasino acquired a PhD in Chemical Engineering at the University of Colorado. At CU, she developed synthetic phospholipids as part of an artificial cell project and acquired experience in microfluidics fabrication. Combining these experiences, she led development of a novel polycarbonate-based organ-on-chip platform as a postdoc at Oregon Health and Science University. Now her lab, the μMicrobiome Lab, aims to further develop and apply gut microbiome-on-chips to study the relationship between the human gut microbiome, distal disease progression, and therapeutic efficacy.

“Genetic and Microbiome Control of Addiction-like Behaviors in Mice”, Dr. Jason Bubier, Ph.D., Senior Research Scientist, Center for Addiction Biology, The Jackson Laboratory
Jason A. Bubier, Ph.D., is a Senior Research Scientist at The Jackson Laboratory, where he leads research at the intersection of genetics, behavior, and addiction biology. His work focuses on uncovering the genetic and physiological factors that shape vulnerability to addictive substances, with current projects examining the microbiome’s influence on behavior, the genetic architecture of opioid‑induced respiratory depression, and systems‑level pathways involved in addiction.
Dr. Bubier has spent more than two decades at The Jackson Laboratory, advancing through multiple research roles and contributing to major initiatives in functional genomics and behavioral genetics. He has authored more than 80 scientific publications, and his work is widely cited across fields including immunology, bioinformatics, and addiction science.

“Food insecurity, gut microbiome, health”, Dr. Maria Carlota Dao, Ph.D., Assistant Professor of Human Nutrition, University of New Hampshire.
Dr. Maria Carlota Dao is an Assistant Professor of Human Nutrition in the Department of Agriculture, Nutrition, and Food Systems at the University of New Hampshire. As an interdisciplinary scientist focused on obesity research, she investigates the interplay of cardiometabolic risks, dietary and psychosocial factors, and the gut microbiota. Through this work she seeks to address obesity in health disparity populations. Prior to becoming a faculty member at UNH, Dr. Dao worked as a scientist at the Jean Mayer USDA Human Nutrition Research Center on Aging at Tufts University. Her training includes a postdoctoral fellowship at the Sorbonne University and INSERM (Institute of Cardiometabolism and Nutrition) in France, and a PhD in Biochemical and Molecular Nutrition from the Friedman School of Nutrition Science and Policy at Tufts University.

“Gut Anthro“, Dr. Amber Benezra, Assistant Professor of Science and Technology Studies at Stevens Institute of Technology
Dr. Benezra is a sociocultural anthropologist researching how studies of the human microbiome intersect with biomedical ethics, public health/technological infrastructures, and care. In partnership with human microbial ecologists, she has been developing an “anthropology of microbes” to address global health problems across disciplines. Her book, Gut Anthro (published by University of Minnesota Press in 2023) is the first ethnography of the microbiome.

12:00 – 1:15 Lunch, provided
1:15 – 2:45 Session 2: Biochemical geniuses: microbes from soil to sea
“How to build a microbiome to “breed” better plant-inoculants.” Dr. Anna O’Brien, PhD., Assistant Professor of Molecular, Cellular, and Biomedical Sciences, University of New Hampshire.
She is originally from the Pacific Northwest. She received her B.S. in Plant Biology from the University of Washington, her PhD from the University of California Davis, where she worked with Jeffrey Ross-Ibarra and Sharon Strauss on the impacts of rhizosphere biota on trait divergence and adaptation in the wild relatives of maize. Anna Post-doc’ed at the University of Toronto, primarily with Megan Frederickson, but also with Chelsea Rochman, David Sinton and unofficially, Stephen Wright, in everything to do with duckweeds and duckweed microbiomes (from mutualisms, to ecotoxicology, to engineering better tools, and experimental “evolution” and epigenetics).
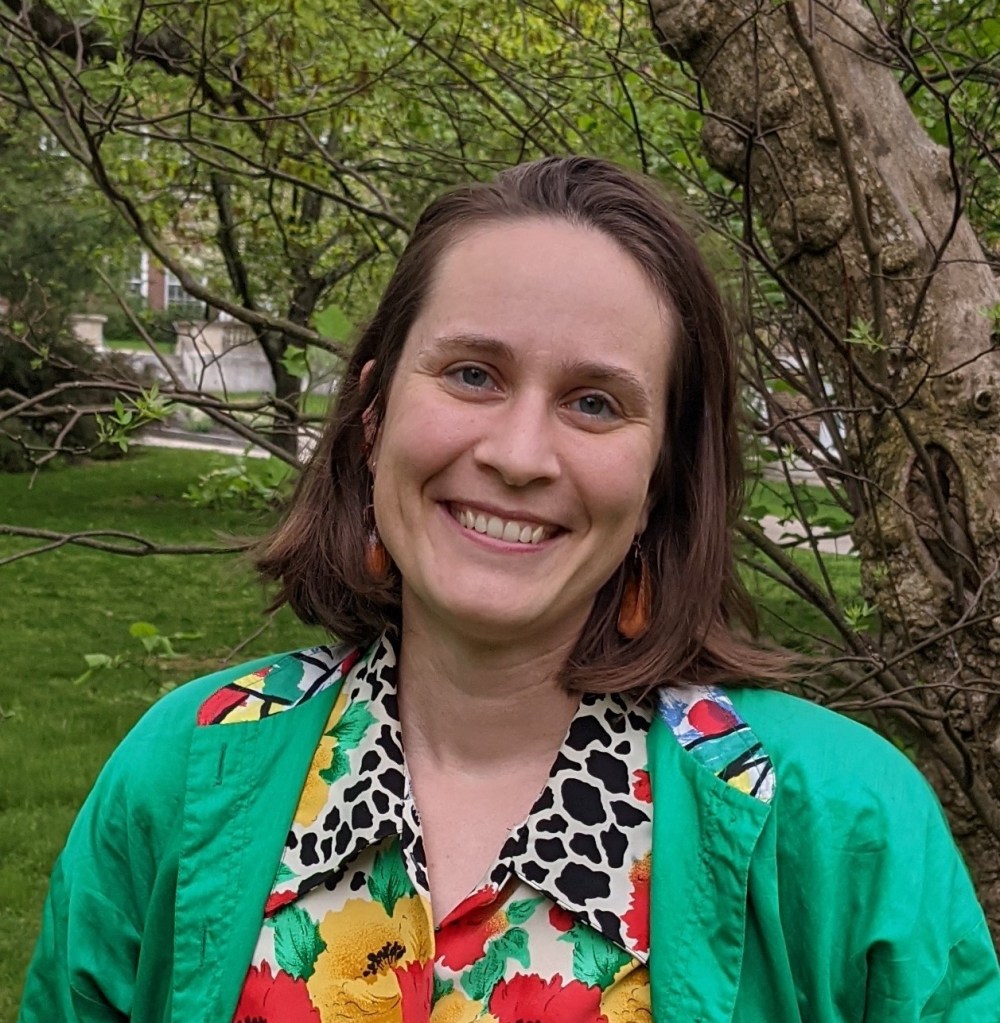

“Salt marsh microbiomes are impacted by waterway restrictions in coastal Maine.” Heather Richard, M.S., PhD. Candidate in Ecology and Environmental Sciences, UMaine.
Heather joined the University of Maine in 2021 as a PhD student with the Maine eDNA program and studies the impacts of bridges and roads on microbial communities in salt marsh habitats. Her background in Ecology led her to pursue a career in informal environmental education for several years before getting a Master’s degree in Marine Biology from San Francisco State University studying biofilms on microplastics pollution. Upon returning to Maine in 2016 she led local research for a coastal non-profit organization and has since been dedicated to studying coastal environmental issues relevant to Maine. She has found a true passion in bioinformatic analysis and is eager to learn new tools for data analysis of all kinds.
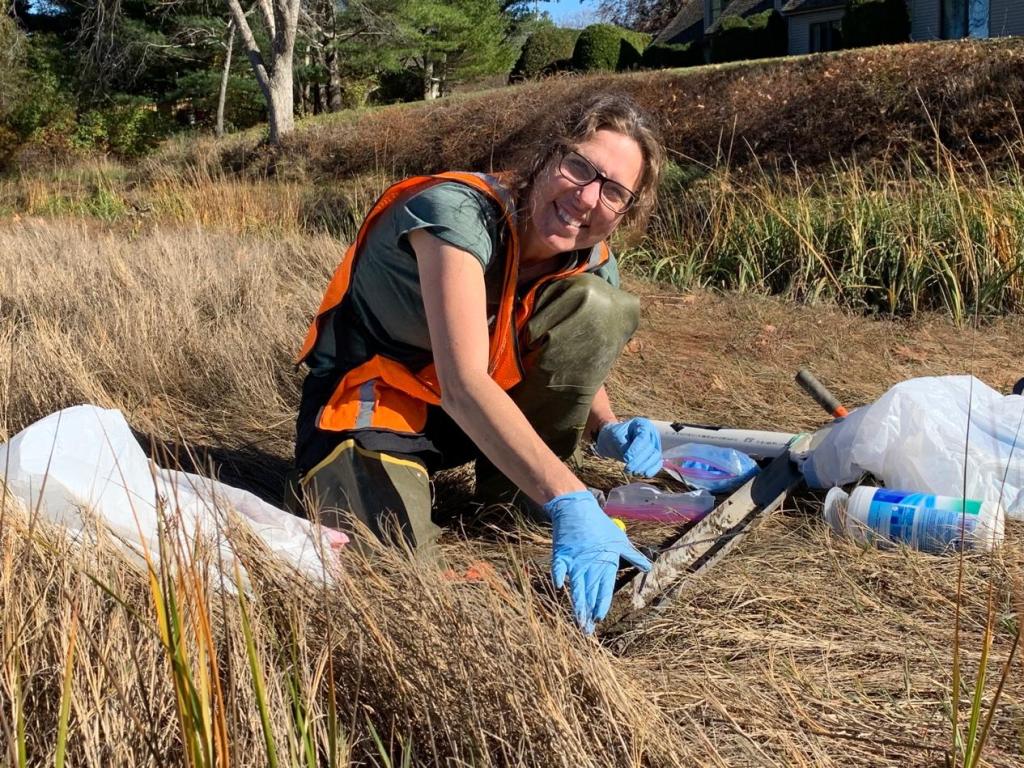

“Genomic Strategies for Discovering Microbial Dark Matter using Metagenomics“ Scott Tighe, M.S., Senior Research Associate, Environmental Microbiome Engineering Research Group, Dept of Civil Engineering, University of Vermont.
Scott Tighe is currently Senior Research Associate in the Environmental Microbiome Engineering Research Group in the Dept of Civil Engineering at the University of Vermont and past technical director of the UVM Genomics Facility. He has expertise in all areas of genomics, microbiology, and mycology with a specific focus on advancing the methods used for microbiome and metagenomic analysis. He specializes performing metagenomic analysis in global extreme sites such Greenland, Antarctica, Romania, Saudi Arabia, Crete, Tahiti, and the International Space Station to name a few and is currently the science lead of the Extreme Microbiome Project (XMP). Short Abstract: The ability to perform advanced genomic techniques on samples collected from extreme environments demands high performance reagents and techniques not commonly used in most labs today. For the past15 years, our projects have developed novel sampling and DNA sequencing protocols for profiling a variety of environments, including snow, glaciers, Antarctic lakes, Romanian caves, thermophilic ecosystems, halophilic, acidic, alkaline lakes, anaerobic digestors as well as organisms from bioaerosols and the International Space Station. Technical approaches include novel liquid concentrating devices, sample disruptors, biopulverizers, and field use of the Oxford Nanopore DNA sequencer. Whole Genome sequencing results indicate that for many unique sights, 5-30% of all organisms are not classified beyond Genus with a large number being new species.
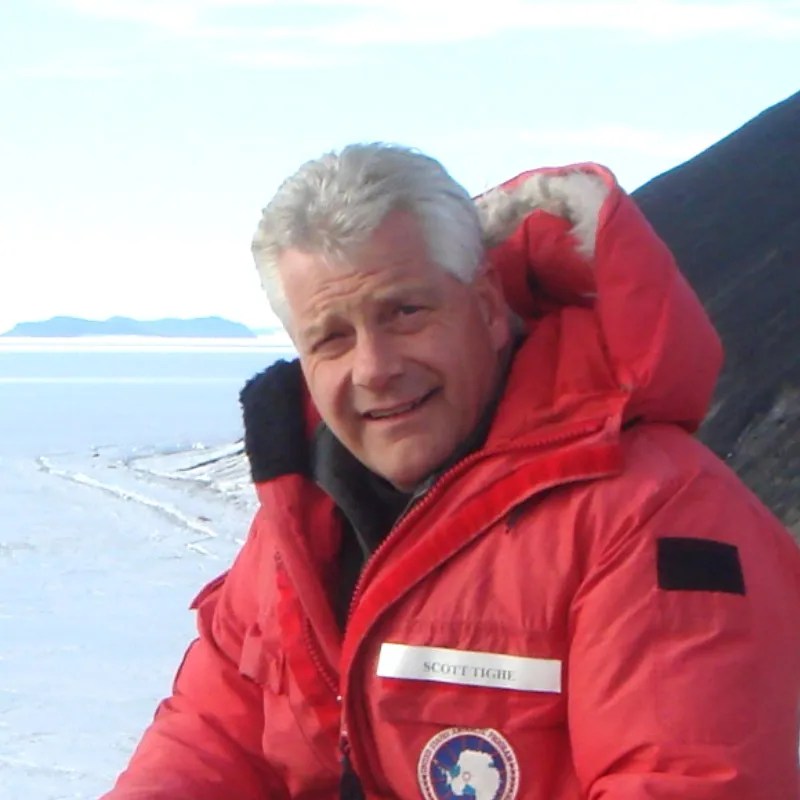

“A Sample-to-results Workflow for Sequencing from Wastewater“, Dr. Siva Chavadi, PhD. Senior FAS, NEBNext Products, NEB
Speaker Bio: Siva Chavadi (he/him) is a Senior FAS (NEBNext NGS sample prep reagents) at New England Biolabs, working with academic and core facilities as well as industrial collaborators to implement scalable NGS workflows. Previously at GENEWIZ/Azenta, he supported NGS service operations and customer applications. He holds a PhD in Animal Sciences from University of Hyderabad, India with research focused on TB diagnostics and vaccine development using functional genomics tools.

2:45 – 3:00 Break
3:00 – 4:00 Session 3: Practical Applications
“Antibiotic-Free Liquid Surface Coatings for the Tunable Control of Protein and Microorganism Adhesion: The rise of antibiotic resistance is one of the greatest global public health challenges of our time”, Dr. Caitlin Howell, PhD, Associate Professor of Bioengineering, Department of Chemical and Biomedical Engineering, University of Maine
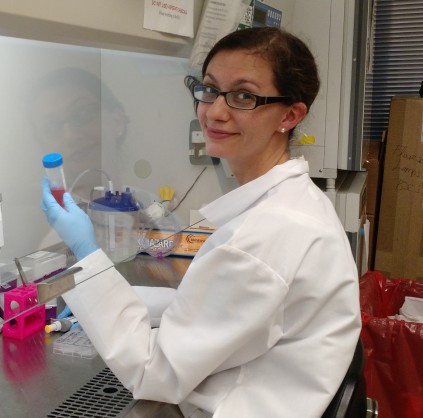

“Beverage safety”, Sam White, M.S., Quality Control Collaboratory (QC2), University of Southern Maine
Samantha White is a dedicated researcher and educator with extensive experience in analytical chemistry and the craft beverage industry. Sam manages the Quality Control Collaboratory at USM where she leads lab operations, quality control initiatives, research projects, and industry outreach programs.

“From Bugs to Breakthroughs: Sequence the Invisible Majority with QIAseq.”, Dr. Kevin W. Joseph, PhD, Genomics Field Application Scientist
Kevin graduated from University of Maine in 2012 with a BS in Microbiology and later acquired his PhD in Infectious Disease and Microbiology from the University of Pittsburgh. During his graduate work, he focused on HIV-1 single genome sequencing and integration site analysis of clonally-expanded, replication competent, HIV-1 proviruses in patients with long-term viral suppression. He developed the Individual Proviral Sequencing Assay (IPSA) to study these rare targets, which enables sequencing of full-length HIV-1 single-genome proviruses and their integration sites with high sequencing efficiency. Kevin has brought his knowledge and expertise to QIAGEN where he assists researchers with NGS, PCR assay and oligonucleotide design, and in-lab training/support.

“Illumina innovation roadmap“, Jennifer Mosher, Senior Sequencing Specialist, Illumina
Jennifer Mosher is a Senior Sequencing Specialist at Illumina, with deep expertise spanning field applications, marketing, and NGS-based proteomics. With over a decade in academia, she previously led operations for NGS at the Cornell Genomics Core before transitioning to industry, where she partners with both academic and commercial leaders to drive adoption of advanced sequencing technologies and assays.

4:00 – 5:00 pm Posters and networking
Agenda for June 12, 2026
This workshop broadly includes combining microbiome data with other data types, combining qualitative and quantitative data, learning to evaluate microbiome data using demographic data, dealing with unbalanced groups especially in human microbiomes, and how to choose statistical tests for microbiome projects.
8:00 – 9:00 am Breakfast, provided
9 – 10- ish am Case Study Presentations, 20 min each, hosted by Sue Ishaq
“Human data pitfalls TBD” Kevin Roberge, PhD. UMaine: Lecturer, Mathematics and Adjunct Professor, Women’s, Gender & Sexuality Studies.
“Statstical decisions TBD” Laura Jackson, Ph.D., UMaine: Bioinformatics Program Coordinator & Graduate FacultyGraduate School of Biomedical Science & Engineering, Bioinformatician – CORE Strategic Operational Services.
“A case study in human microbiomes TBD”, Lola Holcomb, PhD, University of New England, Bioinformatics Postdoc
10 – 10:15 Break, snacks and coffee provided
10:15 – 11 Panel discussion, symposium speakers are welcome to join
11 – 12 Open working time for data wrangling advice and analysis workflow storyboarding as a group
- Kevin Roberge, can help with analysis storyboarding, identifying assumptions, math
- Laura Jackson, can help with coding, statistics and deciding which to use, RNAseq data, 16S data, meta-analyses
- Lola Holcomb, can help with coding, metagenomics data, 16S data, meta-analyses
- Sue Ishaq, can help with some 16S, but will be helping to coordinate the workshop
12 – 1:30 lunch, provided
We are grateful to our financial supporters, who helped make this symposium possible and accessible!
Future Microbiome Researcher Sponsors
These incredible sponsors not only helped support this symposium, but their generous donation helped to create several hands-on laboratory workshops to support training in microbiome technology!






Gold Sponsors
Silver Sponsors


