A collaborative paper on lobster shell bacteria has just been published in the journal iScience: “Water temperature and disease alters bacterial diversity and cultivability from American Lobster (Homarus americanus) shells.” This paper investigates what happens to bacterial communities on healthy and sick lobsters as they experience different water temperatures for a year.
You can read the paper here.


I joined this project back in the summer of 2020, towards the end of my first year at UMaine, when I was given a large 16S rRNA gene sequence dataset of bacterial communities from the shells of lobsters. I had been asking around for data as a training opportunity for Grace Lee, who at the time was an undergraduate at Bowdoin College participating in the abruptly cancelled summer Research Experience for Undergrads program at UMaine in summer 2020. Instead, Grace joined my lab as a remote research assistant and we worked through the data analysis over the summer and fall. Grace has since graduated with her Bachelor’s of Science in Neuroscience, obtained a Master’s of Science at Bowdoin, and is currently a researcher at Boston Children’s Hospital while she is applying to medical school.
My first point of contact on the project was Jean MacRae, an Associate Professor of Civil and Environmental Engineering at UMaine, who was the one to lend me the data and who had been working on bacterial community sequencing on other projects which I’ve been involved in. Jean has been involved with MSE, and this is our fourth publication together making her the collaborator at UMaine I have co-authored with the most (although it is a tight race 🙂 ).


Jean introduced me to the original research team, including Debbie Bouchard, who is the Director of the Aquaculture Research Institute and was researching epizootic shell disease in lobsters for her PhD dissertation several years ago; Heather Hamlin, Professor and Director of the School of Marine Sciences; Scarlett Tudor (not pictured), the Education and Outreach Coordinator at the ARI; and Sarah Turner (not pictured), Scientific Research Specialist at ARI. The ARI team is involved in a lot of large-scale aquaculture research, education, and outreach to the industry here in Maine, and the collaborative work I have been doing with them has been a new an engaging avenue of scientific study for me.
In 2022, the research team, along with social science Masters student Joelle Kilchenmann, published a perspective/hypothesis piece which explored unanswered questions about how the movement of microbes, lobsters, and climate could affect the spread of epizootic shell disease in lobsters off the coast of Maine. That perspective paper was a fun exercise in hypothesis generation and asking ‘what if’?

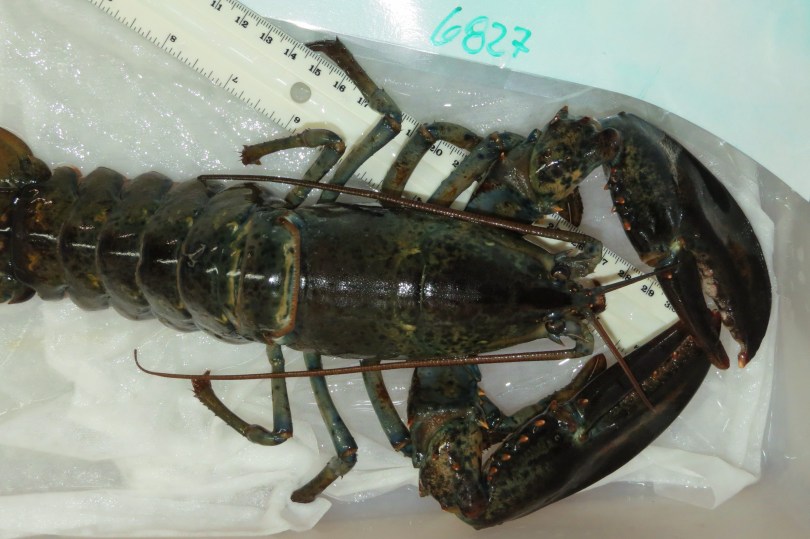
This manuscript is more grounded, and features work that was started in 2016. It examines bacterial communities on the shells of lobsters which were captured off the coast of Southern Maine and maintained in aquarium tanks for over a year. The lobsters were split into three treatment groups: those which were kept in water temperatures that mimicked what they would experience in Southern Maine, colder water to simulated what they would experience in Northern Maine, and hotter water to simulate what they would experience in Southern New England over that year. The original project team wanted to know if temperatures would make a different to their health or microbial communities.

The original project team swabbed lobster shells to obtain bacteria to try and grow in the lab, as well as DNA to sequence and identify whole bacterial communities. Grace and I performed the data analysis to identify which taxa were present in those communities, what happened over time or when the water temperature changed, and what bacteria were present or not in lobsters which died during the study.
Figure S11. Lobster carapace sampling using a sterile cotton swab to obtain bacterial communities from the shell surface, related to STAR Methods. The right side of the dorsolateral area of the cephalothorax was sampled for the baseline sampling, the left side for the Time 1, and the right side again for Time 2.

In addition to wanting to know about temperature, we wanted to know specifically how temperature would affect the bacteria if the lobsters had epizootic shell disease. It is not known what causes epizootic shell disease (which is why it is called ‘epizootic’), but it manifests as pitting in the shells of lobsters. Over time, the pitting can weaken shells and make it difficult for the lobster to molt, or make the lobster susceptible to predators or microbial infections. This type of shell disease had been a huge problem in Southern New England over the past few decades, and in Maine we have seen more cases over time.

The highlights of this project are here, but you can click the link below to read the entire study and what happened to lobster health and lobster microbes over time.
- Shell bacteria from healthy lobsters, often overlooked, were included in the study.
- Hotter and colder water temperatures affected shell bacterial communities.
- Epizootic shell disease reduced bacterial diversity on lobster shells.
- Epizootic shell disease could be induced or exacerbated by the loss of commensal bacteria from shells.
“Water temperature and disease alters bacterial diversity and cultivability from American Lobster (Homarus americanus) shells.”
Suzanne L. Ishaq1,2,, Sarah M. Turner2,3, Grace Lee4,5,M. Scarlett Tudor2,3, Jean D. MacRae6, Heather Hamlin2,7, Deborah Bouchard2,3
- 1 School of Food and Agriculture; University of Maine; Orono, Maine, 04469; USA.
- 2 Aquaculture Research Institute; University of Maine; Orono, Maine, 04469; USA.
- 3 Cooperative Extension; University of Maine; Orono, Maine, 04469; USA.
- 4 Department of Neuroscience, Bowdoin College, Brunswick, ME 04011; USA.
- 5 Boston Children’s Hospital, Boston, MA 02115; USA.
- 6 Department of Civil and Environmental Engineering; University of Maine; Orono, Maine, 04469; USA.
- 7 School of Marine Sciences; University of Maine; Orono, Maine, 04469; USA.
Summary
The American lobster, Homarus americanus, is an economically valuable and ecologically important crustacean along the North Atlantic coast of North America. Populations in southern locations have declined in recent decades due to increasing ocean temperatures and disease, and these circumstances are progressing northward. We monitored 57 adult female lobsters, healthy and shell-diseased, under three seasonal temperature cycles for a year, to track shell bacterial communities using culturing and 16S rRNA gene sequencing, progression of ESD using visual assessment, and antimicrobial activity of hemolymph. The richness of bacterial taxa present, evenness of abundance, and community similarity between lobsters was affected by water temperature at the time of sampling, water temperature over time based on seasonal temperature regimes, shell disease severity, and molt stage. Several bacteria were prevalent on healthy lobster shells but missing or less abundant on diseased shells, although some bacteria were found on all shells regardless of health status.