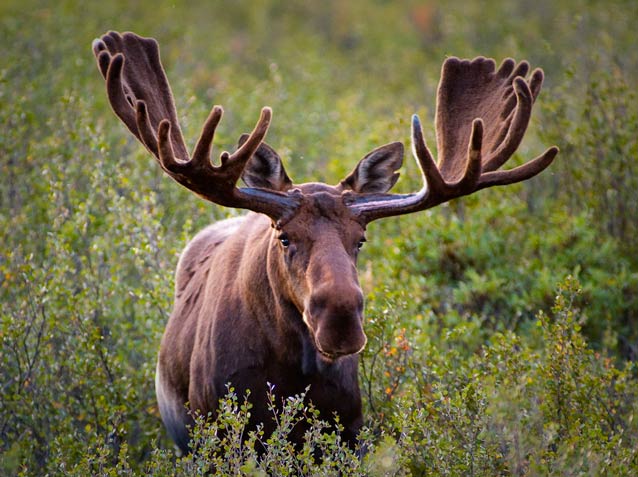
Which and how many bacteria, archaea, fungi, viruses, or protozoa (collectively, the microbiome) might inhabit the tissues or surfaces of an animal host is impacted by myriad factors, including the animal host’s biology, ecology, lifestage, health status, and diet. Much of my lab focuses on understanding the players and the dynamics in the microbial universe that is the digestive tract of various animal species.
The digestive tract of animals is a veritable universe for microorganisms: some pass through but once, some are frequent tourists, and some spend their entire existence in the confines of the gut. Each microorganism brings with it an array of capabilities for digesting or producing different chemical compounds. In many cases, the animal host is reliant on these microbial functions to digest its diet and maintain a hesitant truce with the more exuberant members of the gut community. Each organ in the digestive tract creates a unique environment, with variations in temperature, oxygen content, pH, salinity, muscle contractions, animal cells, and animal enzymes. These conditions, and their effect on the animal’s diet as it moves through the gut, combine as selective factors for which and how many microorganisms thrive in any location. This effect on the microbial communities requires us to use spatial resolution in our studies – or sampling multiple locations along the gut. Yet, this is invasive and logistically difficult to do in live animals, and it prevents us from using temporal resolution – or sampling over time to understand the dynamics of that host-associated microbial community.
Related Reviews
- Garcia-Mazcorro, J.F., Ishaq, S.L., Rodriguez-Herrera, M.V., Garcia-Hernandez, C.A., Kawas, J.R., Nagaraja, T.G. 2019. Review: Are there indigenous Saccharomyces in the digestive tract of livestock animal species? Implications for health, nutrition and productivity traits. Animal: 1-9. Impact 2.026. Article.
- St-Pierre, B., Cersosimo, L.M., Ishaq, S.L., Wright, A-D.G. 2015. Toward the identification of methanogenic archaeal groups as targets of methane mitigation in livestock animals. Frontiers in Microbiology 6:776. Impact 4.165. Article.
- Ishaq S.L., Wright A-D.G. 2015. Terrestrial Vertebrate Animal Metagenomics, Wild Ruminants. In: Highlander, SK, Rodriguez-Valera, F, White, BA. (Ed.) Encyclopedia of Metagenomics: SpringerReference. Springer-Verlag Berlin Heidelberg. Article.
- Ishaq, S.L., Wright, A-D.G. 2015. Wild Ruminants. In: Rumen Microbiology – Evolution to Revolution. AK Puniya, R Singh, DN Kamra (eds). Springer India. Pp. 37-45. Article.
Related Projects
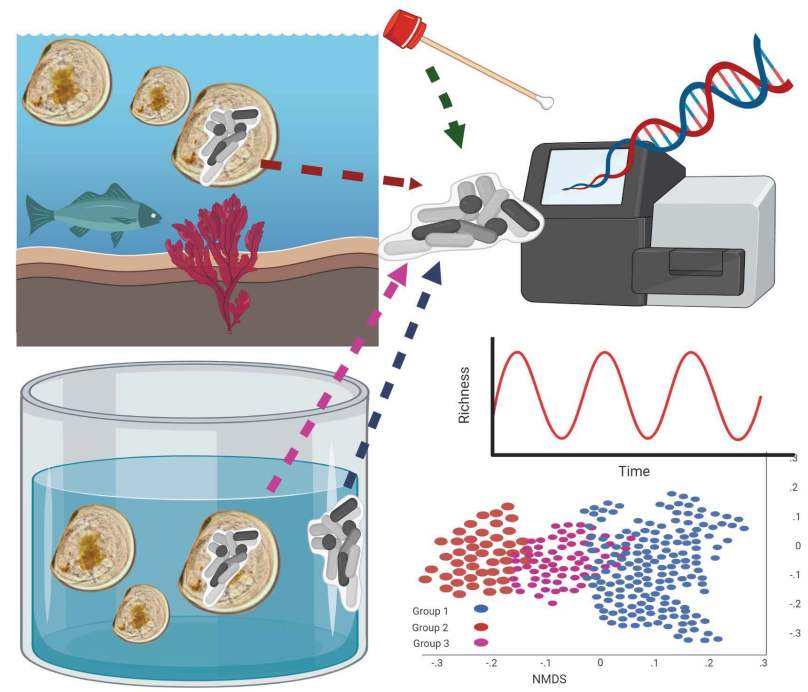
Bacterial community trends associated with sea scallop, Placopecten magellanicus, larvae in a hatchery system.
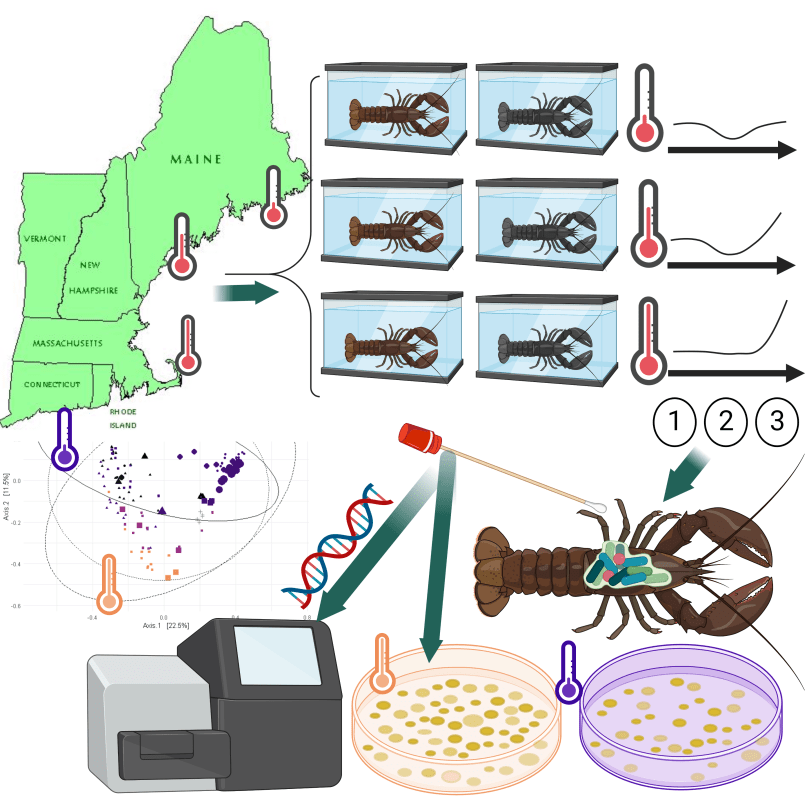
Warmer water temperature and epizootic shell disease reduces diversity but increases cultivability of bacteria on the shells of American Lobster (Homarus americanus).

Many questions remain unanswered about the role of microbial transmission in epizootic shell disease in American lobsters (Homarus americanus).

Determination of the microbial community in the rumen and fecal matter of lactating dairy cows fed on reduced-fat dried distillers grains with solubles.

Bacterial transfer from Pristionchus entomophagus nematodes to the invasive ant Myrmica rubra and the potential for colony mortality in coastal Maine.

Pelleted-hay alfalfa feed increases sheep wether weight gain and rumen bacterial richness over loose-hay alfalfa feed.

Zinc amino acid supplementation alters yearling ram rumen bacterial communities but zinc sulfate supplementation does not.

An investigation into rumen fungal and protozoal diversity in three rumen fractions, during high-fiber or grain-induced sub-acute ruminal acidosis conditions, with or without active dry yeast supplementation.

Biogeographical Differences in the Influence of Maternal Microbial Sources on the Early Successional Development of the Bovine Neonatal Gastrointestinal tract.

Ground Juniperus pinchotii and urea in supplements fed to Rambouillet ewe lambs. Part 2: Ewe lamb rumen microbial communities.

A living soil inoculum increases soil microbial diversity, crop and weed growth using soil from organic and conventional farms in northeastern Montana.

High-throughput DNA sequencing of the moose rumen from different geographical locations reveals a core ruminal methanogenic archaeal diversity and a differential ciliate protozoal diversity.

Fibrolytic bacteria isolated from the rumen of North American moose (Alces alces) and their potential as a probiotic for ruminants.

Dissertation: A Comparative Analysis Of The Moose Rumen Microbiota And The Pursuit Of Improving Fibrolytic Systems




